SEQMOL index

Find 3D neighbors of sequence alignment positions using a PDB file
Download
Once both, sequence alignment and PDB files are provided, it is possible to select an amino acid in the alignment by double-clicking it and then find all neighbors of this amino acid in 3D-space, mapped on the sequence alignment.
These settings control the 3D search:
Example search output: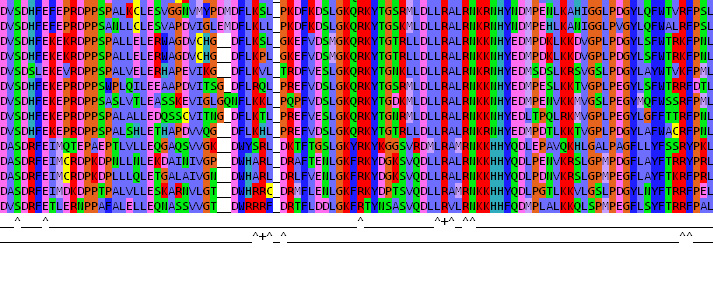
"+" show double-click selected amino acids
^ show partners of this amino acid the PDB file within 5-angstrom sphere
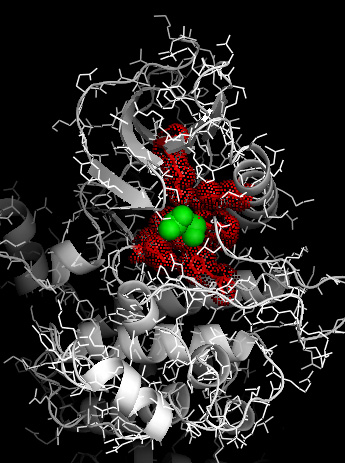
In 3D-space such selections would correspond to a picture like this, where "+" is green residue and "^" are red residues: